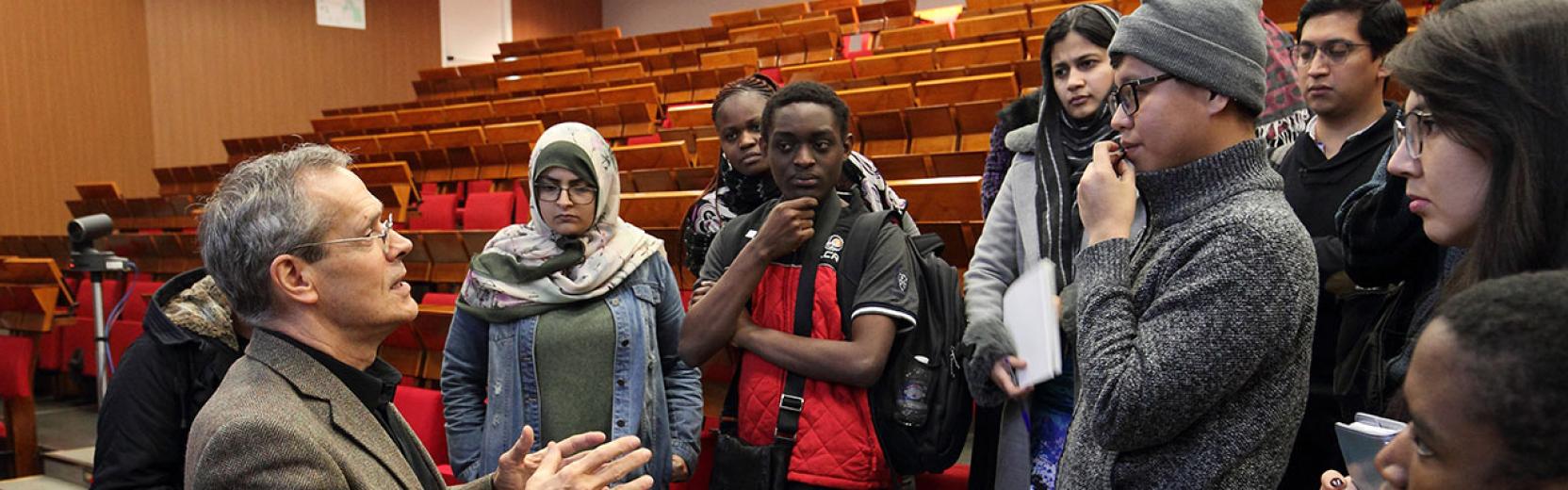
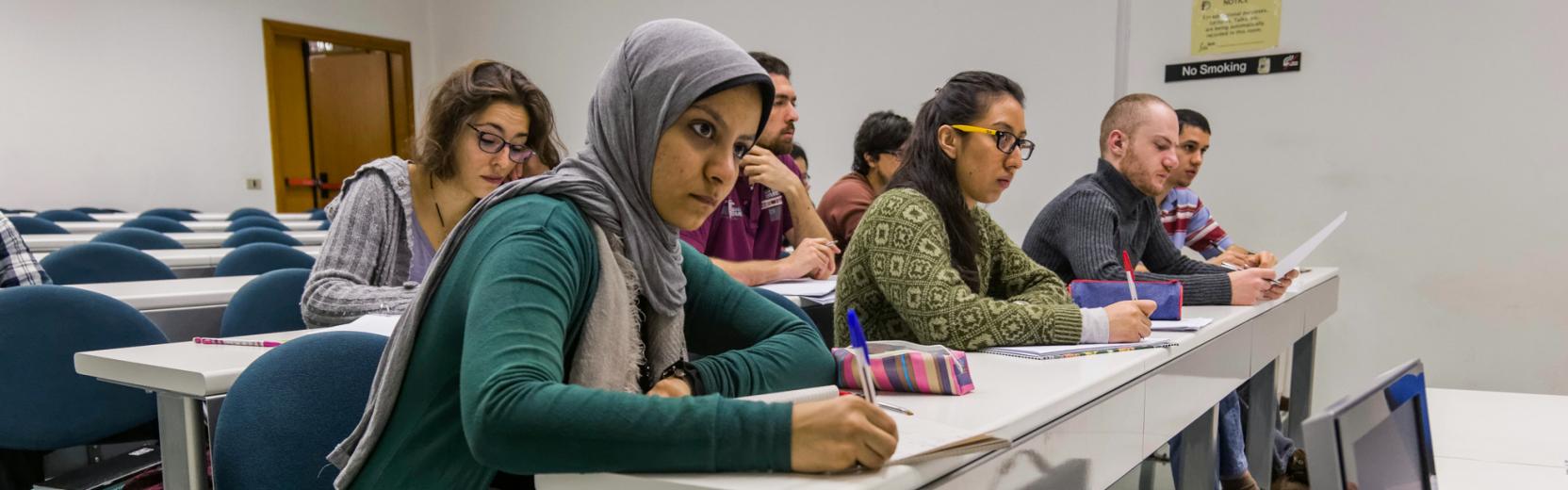
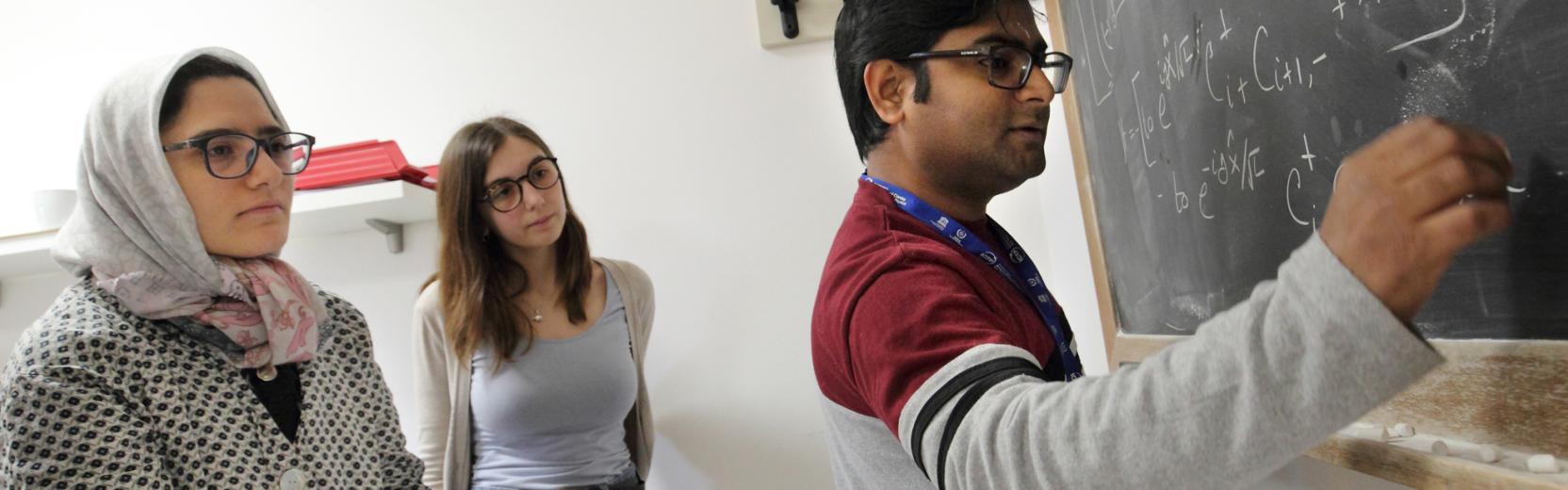
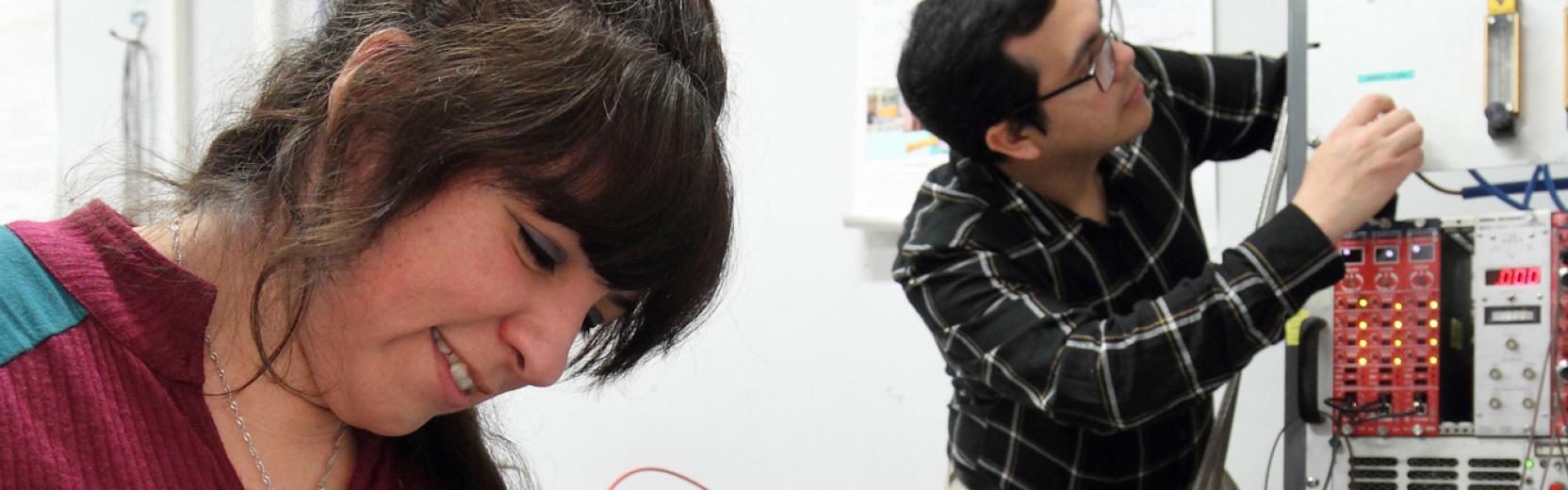
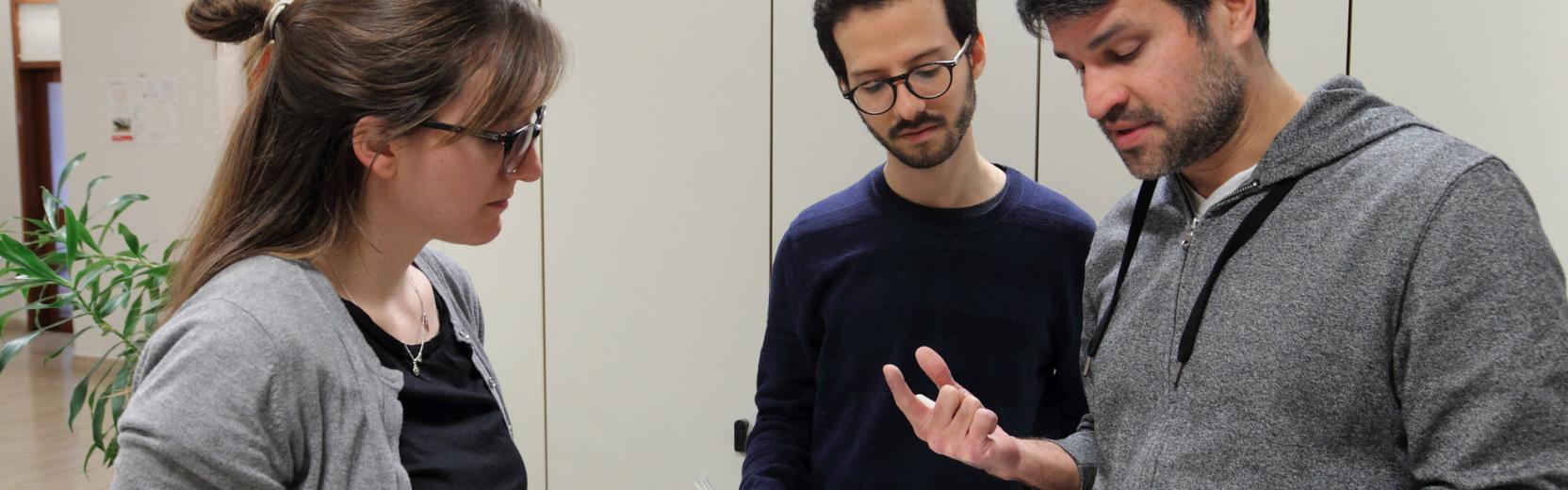
Research in the CMSP Section spans some of the most exciting areas of theoretical condensed matter physics, including the investigation of properties of nanostructures, the study of interacting many-body quantum systems, quantum information processing, computer simulations of fluids and solids with atomistic, molecular, and electronic structure methods, the design of new materials for renewable energy applications and synchrotron-radiation related physics.
Back
Research Areas
Next

Equilibrium and Non-equilibrium Many-body Quantum Systems
The investigations of quantum many-body systems at CMSP bridges different aspects of condensed matter, statistical mechanics and quantum information. The wide spectrum of topics covered includes the study of superconductivity and magnetism in strongly-correlated systems, superfluids, cold atoms in optical lattices, localization in disordered systems, many-body physics with light, dynamics and relaxation in complex systems, quantum simulators, and quantum computing.
One of the most promising routes to building a quantum computer is the engineering of quantum annealers, or adiabatic quantum computers. While some prototypes are already being commercialized, several important theoretical and practical problems are in need of solutions. One of these is an analysis of how the adiabatic algorithm performs on classical optimization problems. Another problem is, how resilient the adiabatic computer's performance is, in the face of influence from the external environment. At CMSP we are actively working on these questions as well as in several other topics related to quantum annealing.
The goal of this research direction is to understand the fundamental physics of Fermi- and non-Fermi liquids, non-conventional superconductivity of cuprates and iron pnictides, physics of heavy-fermion compounds, properties of diluted and dense Kondo lattices, interplay between superconductivity and magnetism in strongly-correlated materials, properties of spin chains and ladders, and exotic magnetism emerging in quantum liquids. All these topics are currently under scrutiny by several members of the CMSP section.
One of the main assumptions of statistical mechanics is that when an isolated system evolves under its Hamiltonian dynamics, it will reach a state of equilibrium where a statistical description holds. Recently, this assumption has been challenged by a series of papers demonstrating that in disordered, interacting systems, equilibration is hindered by quantum effects. This phenomenon has been dubbed Many-Body Localization (MBL) and is the subject of intense research at CMSP.
As Feynman stated in a celebrated paper, the best way to simulate a quantum system is to use another quantum system. Experimental advances in last decades, with cold atoms, trapped ions, etc, have made it possible to study the equilibrium and non-equilibrium properties of strongly correlated systems. This has revived a number of challenging theoretical questions, and we are actively working on this topic in a number of different directions at ICTP:
- the ground state phase diagram of various strongly correlated systems;
- the dynamics of many-body systems from the case of sudden perturbation to the (quasi-) adiabatic regime;
- quantum control of many body systems;
- the entanglement in complex quantum systems.
The properties of open quantum many-body systems were an almost unexplored area until just a few years ago. The steady state phase diagram becomes incredibly rich, displaying a variety of phenomena (novel phases and forms of order not present in equilibrium.; the universality class of the transitions may also be modified. Intense theoretical activity in this direction, has uncovered a number of unexpected features of collective behavior (including quantum synchronization) in open quantum systems. Physical systems where these phenomena can be studied include: exciton-polariton systems, arrays of coupled cavities, and Rydberg atoms. We are interested in exploring critical phenomena and exotic phases in driven dissipative systems, as well as in proposing viable setups for experimental realizations.

Atomistic, Molecular, and Electronic Structure Simulations
When protein folding goes bad, aggregated structures of biological matter known as Amyloid plaques can form. These proteins have been implicated in neurodegenerative diseases such as Alzheimers and have rather unusual optical properties such as being able to fluoresce. An active area of ongoing research is to use a combination of both ground and excited state electronic structure calculations to understand the electronic and optical properties of these systems. Classical molecular dynamics combined with enhanced sampling methods like umbrella sampling and metadynamics are used to explore the conformational landscape of these proteins. These systems also provide good model systems to understand classical problems with bio-physical chemistry such as hydrophobic effects.
Water is the most ubiquitous solvent for biological matter such as proteins and DNA – it acts a lubricant and maintains the structural integrity of biomolecules. Research in this area is aimed at understanding the intimate coupling between the fundamental interactions of water and the biological systems. The physical chemistry of how organic molecules such amino acids and DNA bases interact with water is at the heart of hydrophobic solvation which drives many processes such as protein folding, drug-ligand binding and protein aggregation. A combination of both computational simulations and statistical mechanical theory is used to examine how these solutes perturb the structural, dynamical, electronic and spectroscopic properties of water.
Multiferroic materials exhibit more than one of the so-called ferroic orders, namely magnetic, ferroelectric, and ferroelastic order. Current research is strongly focusing of magnetoelectric multiferroics that are simultaneously ferromagnetic and ferroelectric, and in which an electric field can be used to modify the magnetic properties. Theoretical predictions of the existence of such materials were substantiated in past years by the growth of epitaxial films of a variety of complex and nanostructured oxides exhibiting magnetoelectric properties. Besides scientific interest in their physical properties, such systems have potential applications in a number of devices exploiting electric-field control of magnetization or magnetic properties, including actuators, switches, and new types of electronic memory, spintronic and high-frequency magnetic devices. Using first-principles density-functional calculations, we investigate the microscopic mechanisms generating magnetoelectric effects in multiferroic layered materials and interface systems and explore ways to chemically and structurally tailor the magnetoelectric couplings for targeted electrically controlled magnetic properties.
Graphene is the most widely researched two-dimensional (2D) material. Its many uncommon properties include a semi-metallic character, high electric and thermal conductivity, and unique mechanical properties: graphene membranes have emerged as the strongest materials measured. Together with the high fexibility provided by the nature of sp2 carbon bond, these outstanding properties make graphene films ideal candidates for ultra-thin, ultra strong, impermeable membranes separating different environments. Graphene films can be grown on metal surfaces on which they form a variety of nanostructured physisorbed and/or chemisorbed phases and are able to trap mesoscopic volumes of gas in nanobubbles, demonstrating suitability for application such as gas-storage, or anvil cells for high-pressure reactions inaccessible under ambient conditions. In addition, the large resulting lattice deformation permits to strain-engineer the local electronic and magnetic properties of the film. We presently investigate by means ab-initio computations the formation and properties of nanostructured phases of graphene on metal surfaces and the physics governing the formation and growth of intercalated gas nanobubbles in graphene; this includes in particular rare-gas solidification in nanobubbles at ambient temperature and their growth/ripening with increasing annealing temperature.
Most condensed matter ceases to exist in the form we know it at ambient conditions and transforms into something very different once brought to the highest pressures achievable today in the laboratory or found in the interiors of planets. Liquids solidify, crystalline solids repeatedly change their lattice structure, compounds often decompose, insulators generally turn to metals, magnetism fades away, etc. Understanding, explaining, predicting this evolution under pressure constitutes an important physics challenge as well as a potential opportunity to create new metastable but useful materials. Recent themes include the formation of non-molecular CO2 phases, the mixing behavior of methane/water fluids at planetary conditions, the role of anharmonicity in LiH, the computational discovery of a new spin liquid phase in O2, and new phenomena in high pressure frictional shear.
Atomistic methods based on the ab-initio description of the electrons guarantee a chemically accurate description of the forces at play but their computational load grows rapidly with the size of the system.
Accurate force-fields parameterized on ab-initio calculations are a promising way to extend the size and time constraints of ab-initio simulations. Extensive work on polarizable models for oxides has shown that accurate potentials can be constructed that reproduce faithfully the ab-initio results on homogeneous bulk phases. Compounds studied so far include SiO2, MgO, MgSiO3, TiO2, and water. The force-field developed for water, for example, is all-atom, dissociable, and capable of describing with reasonable accuracy also the dissociated state.
One of the most fundamental processes in acid-base chemistry is the ionization of water, a rare event where a water molecule will spontaneously ionize and form a hydronium ion and a hydroxide ion. The microscopic fluctuations that drive this process remains poorly understood. Research in this area is aimed at understanding the molecular mechanisms by which the proton and hydroxide ion diffuse through bulk water systems. How these processes change at hydrophobic interfaces such as the surface of water and protein surfaces and how these properties are related to spectroscopic features of experiments is an active area of research. A particular area of ongoing development is probing the properties of water as a 3D-network which will hopefully help its collective structural and dynamical signatures.

Materials for Renewable Energy Applications
Efficient collection and storage of renewable forms of energy like solar radiation or wind requires the development of advanced functional materials. CMSP research in the field of sustainable energy focuses on this materials-related aspect. Using modern computer simulation techniques, conversion and storage processes are investigated on the atomic scale. The research topics include nanostructured solar cells, battery materials, and photocatalytic water splitting.
The desire to employ intermittent renewable energy sources to produce electricity, as well as the spread of electric vehicles, will increase the importance of electricity storage. Metal-air batteries are particularly interesting for their high energy density, but high overpotentials mean that losses upon charging and discharging are limiting their practicality. To solve this issue, we work to understand the basic electrochemical reactions taking place in these batteries during formation and dissolution of oxides of lithium and sodium, as well as in the presence of electrocatalysts such as MnO_2. Computer simulations using density functional theory are our main tool.
Catalysts are an important ingredient for many reactions and thus for many applications, but they are often not very active and can be very expensive. In order to overcome these limitations, it is necessary to understand their working mechanisms and to create catalytic systems with a high degree of effectiveness. To this end, we are investigating molecules, called phtalocyanines, which are related to heme molecules critical in blood. The center of these active catalysts consists of a single atom, which allows high-precision studies of the reaction mechanisms.
Hydrogen gas is an important chemical feedstock for the synthesis of many materials. It can also be used as energy carrier and for energy storage. Thus, the efficient production of oxygen and hydrogen by splitting water molecules is a key process in the quest for sustainable energy sources. Using computer modeling, we are simulating how this reaction takes place on hematite (Fe_2O_3) surfaces. Many properties of this material make it very appealing as anode in photo-electrochemical water splitting reactions, but major aspects of this process remain to be understood.
In dye sensitized solar cells (DSSCs) light is absorbed by molecules (dyes), which are attached to the surface of wide-bandgap semiconductors. For the paradigmatic case of organic molecules on TiO_2 surfaces, we are investigating the interactions of the dyes with the surface, the electronic structure and excited electronic states. Such information, obtained from calculations based on density-functional theory, can help to better understand the stability of DSSCs and the efficiency of the electron transfer, which is at the heart of the functioning of such solar cells.
One of the current challenges for researchers in renewable energy is to find a way to use solar energy to produce fuels, like hydrogen and hydrocarbons, from water and carbon dioxide. This would make the energy cycle carbon-neutral, while retaining the advantage of using liquid and gaseous fuels. We investigate by density functional theory why some titanium-based photocatalysts are more active than pure titania for these reactions. The systems considered are comprised of black hydrogenated titania, sodium titanate and copper clusters at titania surfaces.
Physics of Nanostructures
Understanding the physics of nano-devices offers the fascinating opportunity to explore quantum properties of matter at the mesoscopic level. Besides their fundamental impact on modern technology, quantum coherence and strong interactions play a fundamental role in these systems. CMSP research topics include charge and heat transport, thermoelectric effects, nanoelectromechanical systems, quantum coherent electronics and spintronics, solid-state quantum information processing, and low-dimensional systems with interactions.
Sliding friction and mechanical dissipation, a subject of enormous practical importance in technology, was until recently viewed mostly at the macroscale. Which have turned into a lively microscopic physics playground after the advent of nanoscale tools such as, Atomic Force Microscopes, Surface Force Apparatuses, Quartz Crystal Microbalances, etc. Understanding nanofriction is a challenge because of its non-equilibrium physics. Tackling this challenge can potentially lead to identify the mechanism for its control. Current approaches in the ICTP-SISSA extended group, also sustained by an European Research Council Advanced Grant, move on three fronts. The first is addressing individual problems posed by experiments, mostly by means of classical non-equilibrium molecular dynamics. This area includes the superlubric sliding of clusters, nanotubes, and charge-density-wave systems. The second addresses the role of quantum mechanics in nanofriction, including the building of an ab-initio approach to electronic friction, and quantum phenomena in the sliding of atomic ion chains. The third front, and the most ambitious, is the building of a bona fide theory of sliding friction, something quite promising which just began.
Some of the distinct features of nanostructures appear in their electronic and heat transport mechanisms. Many of these systems have properties that arise from two main ingredients, phase coherence and strong local correlations. CMSP researchers currently study several of these systems and mechanisms. Including transport through quantum dots and wires, hybrid structures, molecular transistors, topological superconductors, spintronic and nanoelectromechanical systems. In nanoelectromechanics both fundamental and applied aspects are investigated. Phase coherent nanostructures are also studied for their important role in newly emerging quantum technologies, such as solid state quantum computation. Ballistic electron transport in nanocontacts through magnetic impurities including adsorbed radical molecules, is studied by combining the many-body physics of the Kondo effect, with parameters obtained from ab initio electronic structure and methods.
Do thermal machines retain their properties when they are scaled down to dimensions where quantum mechanical effects are prominent? This question embraces a number of important aspects of heat and work conversion in small devices, ranging from fundamental problems in determining the ultimate limits, to the mode of operation of thermal machines, to more applied aspects of heat control at the nanoscale. In the CMSP section we are currently interested in those questions and actively studying the properties of quantum heat engines.
Parallel to the development of the theory of quantum information, since many years, there is an ongoing interest in finding physical systems where quantum information processing could be implemented. In an ideal situation, one should identify a suitable set of two-level systems (sufficiently decoupled from any source of decoherence) with some controllable couplings among them, needed to realize single qubit and two-qubit operations. The quest for large scale integrability and flexibility in the design, has promoted nanostructures among the most promising platforms for quantum information processing. Researchers at CMSP are locking into the properties (and protocols) of solid-state qubits, both in semiconducting and superconducting nano-devices.